A rapidly advancing technology („next-generation sequencing“) now allows the acquisition of large-scale DNA sequence or transcriptome data even in non-model organisms. There are at least three reasons why marine genomics is interesting: Marine systems encompass habitats that are unparalleled on earth, for example in the deep sea or in polar areas. Second, life originated in the sea, and hence, the deep phylogenetic diversity of bauplans is highest in the oceans. Third, the role of microbial life including viruses for ecosystem processes and within symbiosis is probably huge, yet the only tool to characterize this diversity are (meta)-genomic tools. One prime goal of EV to apply such techniques is the identification of genetic changes associated with rapid evolutionary adaptation to global change. A related issue are genes and pathways that protect animals against pathogen attack, and in periods of stress such as ocean warming or acidification. Future research will be directed into the combined action of stressors as selection factors.
We use genomic/transcriptomic tools within several projects, among those are (i) comparative sequencing of Syngnathid and related genomes in a study of MHC class II loss [PIs Olivia Roth, Thorsten Reusch, collaboration with Norwegian Sequencing Centre Oslo] (ii) transcriptome sequencing of several deep sea mussel species (Bathymodiolus spp) in order to obtain high-resolution SNP markers to study gene flow, hybridization and speciation [with PhD student Corinna Breusing, Co-PI Frank Melzner] (iii) charaterization of immune genes in comb jellies Ctenophora [PI Thorsten Reusch, collaboration with Olivia Roth, co-PI P.Rosenstiel et al., IKMB Kiel] (iv) rapid transcriptome evolution in sticklebacks as response to trematode infection [Jenny Rieger, David Haase, PI Thorsten Reusch , co-PIs Erich Bornberg-Bauer, U Münster, Martin Kalbe MPI Plön] (v). genetic changes in Emiliania huxleyi upon 1000+generations of adaptive evolution under ocean acidification via reseqeuncing [Kai Lohbeck, Thorsten Reusch, collaboration with P.Rosenstiel et al., IKMB Kiel].
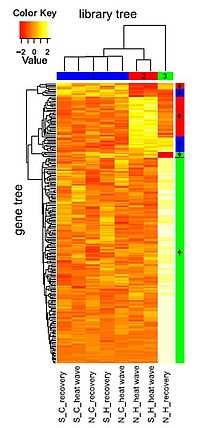
Figure: Heat map illustrating transcriptomic resilience in a marine keystone plant. Franssen et al. PNAS 2011